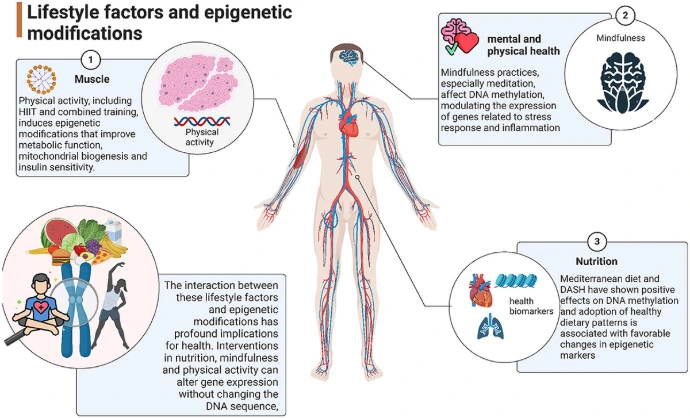
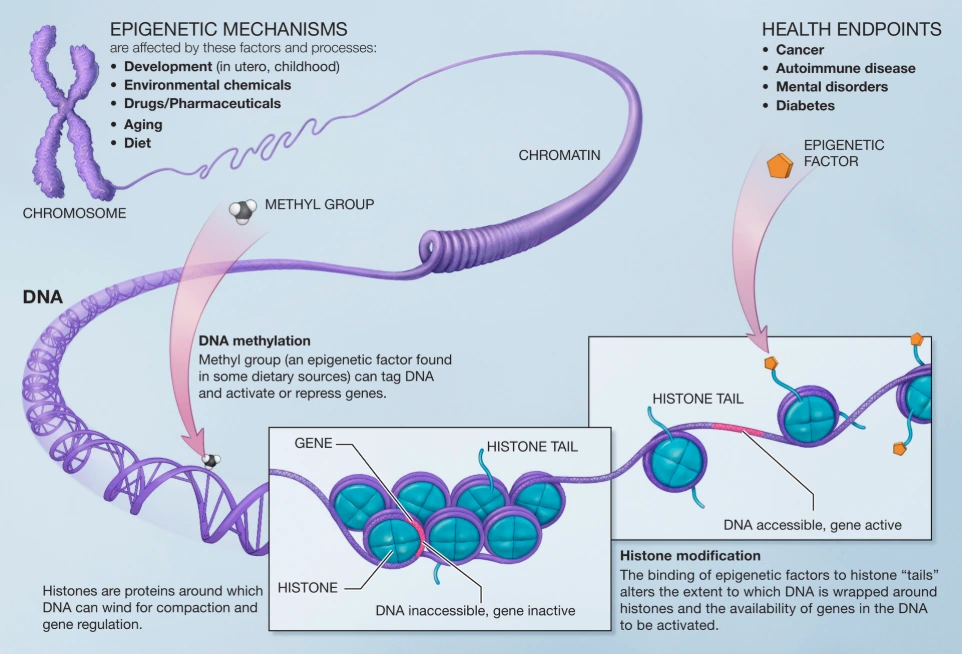
Epigenetic Biomarkers in Diet: Unlocking Precision Nutrition Through Nutrient-Gene Interactions
Introduction
Epigenetic biomarkers represent reversible modifications to gene expression—primarily DNA methylation, histone modifications, and non-coding RNAs—that respond sensitively to dietary inputs. These marks serve as molecular readouts of how nutrients influence metabolic health, inflammation, and aging processes.
Recent comprehensive reviews emphasize that epigenetic changes, especially DNA methylation, explain much of the individual variability in responses to diet and lifestyle, positioning them as promising tools for precision nutrition.
These biomarkers bridge environmental exposures (e.g., folate-rich foods providing methyl groups) to long-term health, enabling tailored interventions for chronic disease prevention.
What Are Epigenetic Biomarkers?
Epigenetic biomarkers are measurable modifications that regulate gene activity without altering the genetic code, reflecting dietary and environmental influences on cellular function.
Key types include:
- DNA methylation (addition of methyl groups to cytosine bases, often at CpG sites, typically repressing gene expression).
- Histone modifications (acetylation, methylation, etc., altering chromatin accessibility).
- Non-coding RNAs (e.g., miRNAs regulating post-transcriptional gene control).
In nutrition, these respond to micronutrients (e.g., B vitamins for one-carbon metabolism) and bioactive compounds (e.g., polyphenols), making them ideal for tracking diet-induced changes.
Tools and Techniques for Discovering Epigenetic Biomarkers in Nutrition
Advanced omics technologies enable genome-wide profiling of diet-responsive epigenetic marks.
Epigenome-wide association studies (EWAS) identify CpG sites associated with dietary patterns or specific nutrients, while epigenetic clocks (e.g., Horvath, PhenoAge, GrimAge, DunedinPACE) quantify biological aging acceleration modulated by diet.
Machine learning integrates multi-omics data (genetics, epigenetics, metabolomics) to predict responses.